Gene Organization
The DNA is a linear string of symbols, A,T,G and C. Proteins are synthesized by reading a code from DNA sequence, with a triplet of nucleotides (a codon) corresponding to a given amino acid. Since 20 amino acids are the constituents of naturally occurring proteins, and there are 64 (= 43) codons, the genetic code is degenerate. The genetic code also includes a rule for initiation of protein synthesis (the start codon) and a rule to signal the end (the three stop or non-sense codons) (Fig. 2.9).
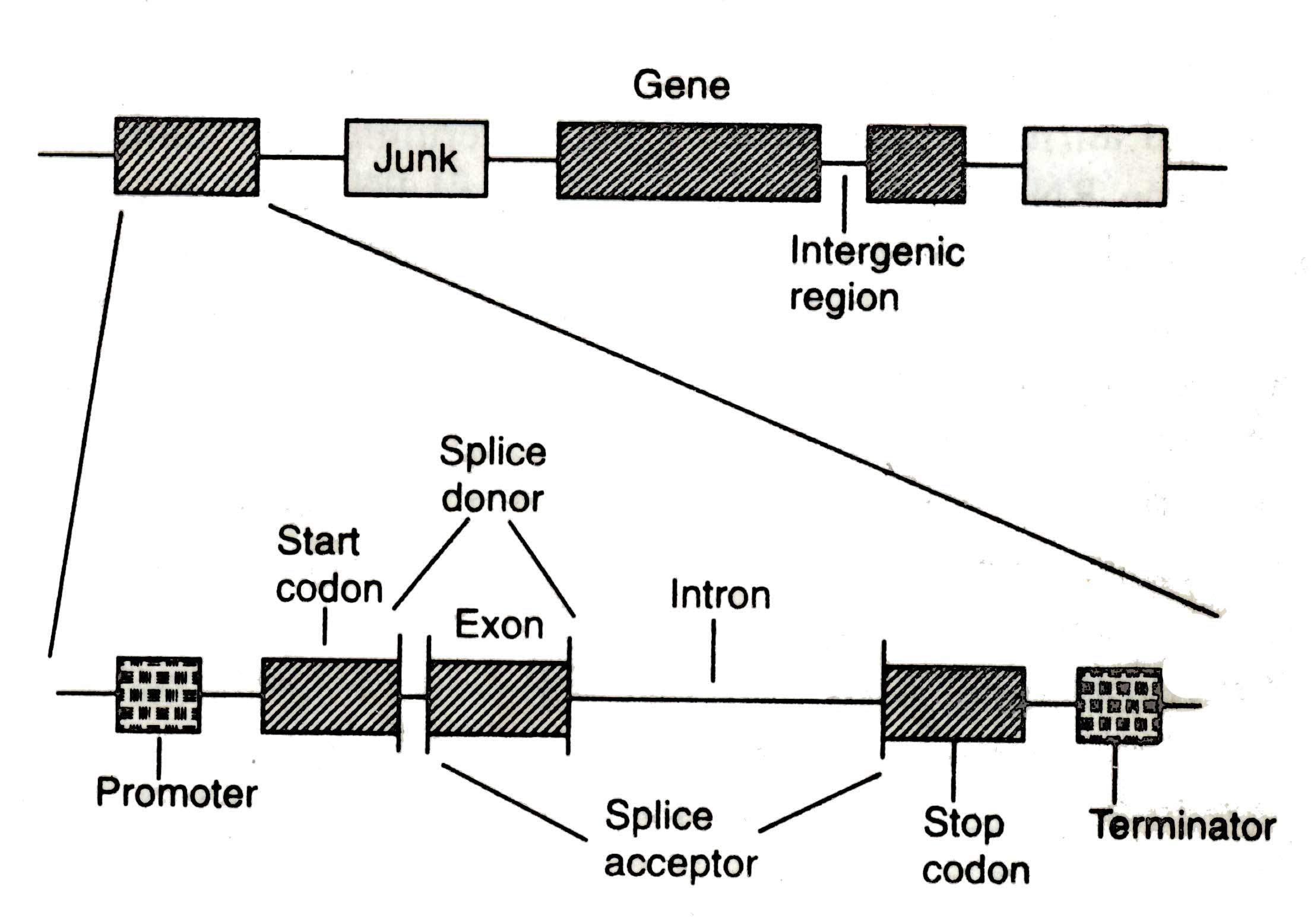
Fig. 2.9. Schematic presentation of different regions in and around a gene in a genomic sequence showing the organization of exons, introns, initiation and termination sites, intergenic spacers and promoters (after Tewari et al., 1996)
The prokaryotic genes are often continuous open reading frames (ORFs) i.e. they are no misprints or interruption, while eukaryotic gene is split into several discrete segments called 'exons' which are interspersed with non-coding intermediate regions i.e. the introns. Exon may be mixed and matched in various combinations to create new genes. Some times exon of one gene may be intron of another gene. The entire gene is transcribed into an RNA molecule, from which introns are spliced out resulting in mRNA. The mRNA is a continuous ORF which is translated into corresponding polypeptide. There are also ancillary regions or the DNA which regulate and control the expression of proteins at specific time and under specific conditions (Tewari et al, 1996).




